 |
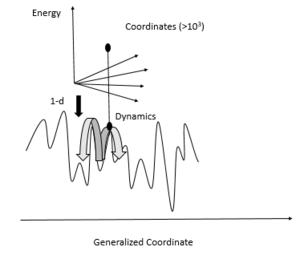
| Fig. 1: Schematic representation of energy landscape (Source: Y. Cao - after Leeson et al. [2]) |
Proteins have four levels of structures: primary, secondary, tertiary and quaternary. The primary structure of protein is a sequence of amino acids which are connected through polypeptide bonds. The polypeptide backbone is then organized into secondary structures such as alpha helices, beta strands and beta sheets. In its tertiary structure, proteins fold into a 3D compact globular structure. The more complicated quaternary structure is an association of tertiary subunits. [1]
Proteins are folded and their structures are maintained by molecular interactions. The balance between protein structural order, i.e. the well-defined tertiary structure, and structural disorder is important to protein function. Protein systems are dynamic and conformational rearrangement of proteins is crucial to their functions. [2] The energy landscape of proteins is often characterized to better understand protein function. The protein energy landscape is the potential energy hypersurface spanned by the relative coordinates of atoms. [2] Protein energy landscapes are usually described in one-dimension as the projection of a single coordinate of the multidimensional conformational phase space, and have a steep-well overall shape, as shown in Fig. 1. [2] Favorable interactions are maximized and unfavorable ones are minimized in the native conformation of the protein, and the low- energy conformations of the protein typically have a rugged structure on the energy landscape. [2] These rugged features correspond to nearly degenerate conformational structures, each of which has slightly different structural details of the protein. [2]
The myoglobin energy landscape has been studied quite extensively. Frauenfelder et al. proposed a model which assumes a hierarchy in the energy landscape. [3] Going down this energy hierarchy, the conformational substate barriers decrease, such that very similar conformations are more readily accessible than ones with larger differences. In addition, the protein structural dynamics have global fluctuations on long timescales and generally only local motions on shorter timescales. [3] Leeson et al. validated the model and obtained a detailed description of this proteins energy landscape by freezing out the large- scale motion. These experiments used optical line narrowing techniques on myoglobin in a temperature range of 0.1-23 °K. [2] Photon echo and spectral hole burning experiments enabled construction of images of the low energy section of myoglobins energy landscape.
Despite the gradual commercialization of catalytic combustion, there are still several challenges. The combustion process itself is highly exothermic with varying fuel/air ratios and fuel composition. The stability of catalyst during the entire operating process is thus a major determining factor of the effectiveness of catalytic combustion. In addition, the wetting of the catalyst surface caused by liquid fuel has to be avoided through robust revaporization technology. [2]
Despite detailed information on the protein energy landscape obtained through experiments, it is still not clear the nature of conformational fluctuations. It is highly advisable to apply similar techniques to other types of proteins in the same and other classes to broaden our current understanding of protein energy landscapes.
© Yi Cao. The author grants permission to copy, distribute and display this work in unaltered form, with attribution to the author, for noncommercial purposes only. All other rights, including commercial rights, are reserved to the author.
[1] T. M. Devlin, Textbook of Biochemistry with Clinical Correlations, 7th Ed. (Wiley, 2010).
[2] D. T. Leeson and D. A. Wiersma, "The Energy Landscape of Myoglobin: An Optical Study," J. Phys. Chem. B 101, 6331 (1997).
[3] H. Frauenfelder, G. A. Petsko and D. Tsernoglou, "Temperature-Dependent X-Ray Diffraction as a Probe of Protein Structural Dynamics," Nature 280, 558 (1979).